Diversity panels (approximately 250 genotypes) of bread and durum wheat and a collection of 24 potato varieties have been defined and assembled. The selection was made by the French National Institute of Agronomic Research (INRA), the National Research Council (CREA) in Italy, ARVALIS, the UK-Based James Hutton Institute and SOLYNTA and was based on a priori genetic and phenotypic information.
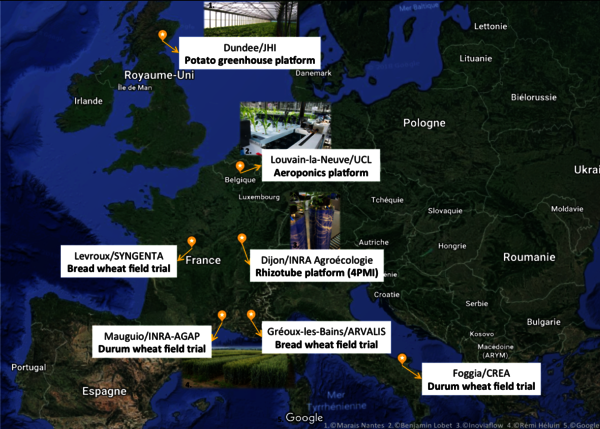
Two field trials were installed for each of the wheat panels during autumn 2017. Bread wheat trials were sown in Levroux (SYNGENTA, France) and Gréoux-les-Bains (ARVALIS, France), and durum wheat trials were sown in Foggia (CREA, Italy) and Mauguio (INRA, France). At all places, each genotype will be exposed to combined water and nitrogen limitations and compared to a control scenario. Field partners are currently working with modellers to establish the final list of variables that will be measured.
The 24 potato varieties have gone through a first greenhouse assay to confirm the interest of the collection. The next trials are in preparation. They will focus on root system architecture, yield and canopy development, and will come up with a novel diagnostic transcriptome test.
We will soon begin the high-throughput root phenotypic platform trials that will be performed at Université catholique de Louvain (Belgium) and at INRA-Dijon (France).
By the end of the summer, the field and platform data from wheat experiments will be analysed together and discussed broadly with partners to determine a short collection of 40 genotypes that will be analysed in more detail within intensified field and platform trials.
More information about work package 2.
 tap and then scroll down to the Add to Home Screen command.
tap and then scroll down to the Add to Home Screen command.